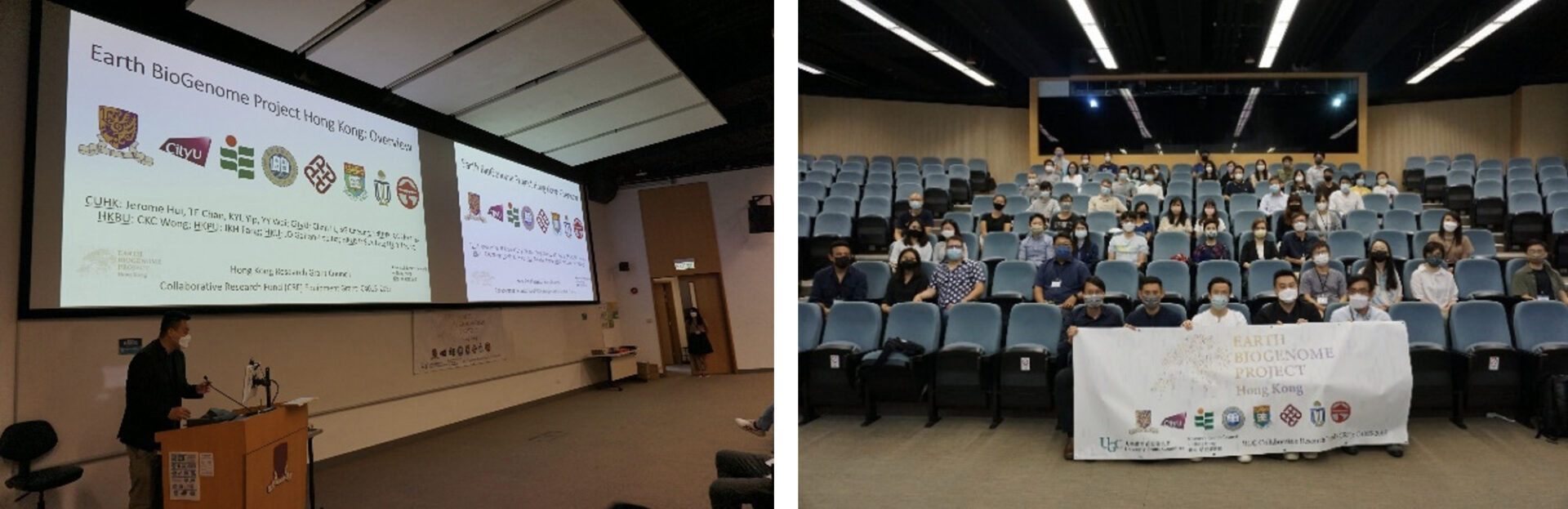
APRU-EBPHK Joint Conference Drives International Collaborations to Slow Biodiversity Loss
Photo Credit: The Chinese University of Hong Kong
“Collaboration” was reiterated at the 2024 APRU x EBPHK Joint International Conference on Biodiversity, Conservation, Genomics and Sustainability from February 15 to 17 at The Chinese University of Hong Kong (CUHK) that attracted 127 participants from 17 universities and other organizations from eight economies.
The three-day conference, jointly organized by the APRU Biodiversity and Sustainability Program and Earth BioGenome Project: Hong Kong and hosted by CUHK, gathered academics from the Asia-Pacific region, as well as NGO practitioners, schoolteachers and members of the public from Hong Kong. Participants shared insights into biodiversity, conservation, genomics, and sustainability, against the backdrop of over a million species being under threat of extinction.
49 speakers delved deeply into a wide range of subtopics, including genetics and genome evolutionary biology research in response to climate change, biodiversity of bacteria, fungi, insects and plants, human-environment interactions for sustainable cities and sustainable lifestyles, as well as biodiversity education.
“Collaboration is key to the goal we wish to achieve, and collaboration cannot stop at the borders,” said Professor Alan Chan, Provost of CUHK.
Prof. Chan, who serves on the APRU Biodiversity and Sustainability Program Steering Committee, added that “we will generate new knowledge that improves our understanding of technological intervention that allows us to better manage the resources that are part of the human heritage.”
The APRU Biodiversity and Sustainability Program was established in 2021 drawing its strength from APRU member institutions representing a significant portion of the world’s research and knowledge capabilities on biodiversity. The program was jointly led by Prof. Jerome Hui from CUHK and Prof. Nathan Lo from The University of Sydney.
Earth BioGenome Project: Hong Kong (EBPHK) is another organizer of the conference. The Hong Kong initiative is derived from the Earth BioGenome Project, which is described as a moonshot project for biology and aims to sequence, catalogue, and analyse the genomes of all eukaryotes on Earth, including animals, fungi, and plants. As a local chapter of the global project, EBPHK was jointly established by eight publicly funded universities in Hong Kong, with an initial focus on organisms that are in high concern and great interest in the city.
The conference was complemented with a field trip to Tai O, a traditional fishing village on Lantau Island that is endowed with a wide variety of natural habitats and species of animals and plants with conservation value.
“At the moment we are losing species at a frequency over a thousand times faster than species were lost before human lived on this planet,” said Professor Kathy Belov, Pro-Vice-Chancellor of The University of Sydney.
Also an APRU Biodiversity Program SC member, Prof. Belov added that “today, you are here to think about how to drive solutions to address biodiversity loss, to better protect the ecosystems, and to combat the impacts of climate change.”
APRU’s Chief Executive Professor Thomas Schneider pointed out that the program provides a platform to support catalytic partnerships and collaborations.
“By connecting scientists, as well as experts and key stakeholders from other sectors of society, the hub creates new incentives for collective action to protect Asia Pacific ecosystems, address biodiversity loss and combat the impacts of climate change,” Prof. Schneider said.
Here is a news article on the conference published by CUHK.
March 12, 2024
more
CUHK Biologists Unveil the Genetic Histories of Centipedes and Millipedes to Contribute to Studies of Biodiversity and Ecology
Pioneering Study of Centipedes and Millipedes Breaks New Ground for Biodiversity
A new genome-sequencing study by CUHK biologists has uncovered the hidden genetic histories that explain differences in the behaviour and diets of centipedes and millipedes. These surprising evolutionary insights could help scientists better understand the vital ecological roles that these creatures play in sustaining and restoring natural ecosystems. The findings were reported in the prestigious scientific journal Nature Communications.
Centipedes prey on insects and other invertebrates, while millipedes feed on leaf litter and other decaying organic matter. Both belong to a group of invertebrate animals called myriapods, which means ‘10,000 feet’ in Ancient Greek and includes about 16,000 extant species. Myriapods perform many crucial ecological functions, including recycling nutrients in the soil and keeping forests healthy. Despite their importance, however, myriapods are relatively understudied compared to invertebrates that share similar features, such as insects, arachnids and crustaceans.
A fork in the family tree
In a major advance for myriapod knowledge, a CUHK team has now sequenced the whole genome of nine centipede and millipede species, creating high-quality reference genomes that constitute the world’s first myriapod gene repertoire analysis. These genomic studies revealed several unexpected gene alterations that led to the different adaptive pathways followed by centipede and millipede lineages after their divergence from a common ancestor, according to Prof. Jerome Hui of CUHK’s School of Life Sciences.
‘This remarkable divergence has led to two very different lifestyles being expressed in extant myriapods: predation in centipedes, characterised by the evolution of offensive chemical weaponry in the form of venom, and a detritivorous diet in millipedes,’ explains Prof. Hui. ‘We provide the first steps towards unravelling the genomic bases of the divergent adaptations underlying these two lineages with very different ecologies.’
Applying genomic insights to promoting biodiversity
Prof. Hui was part of the consortium that published the first centipede genome. He has led a research team working on myriapods since 2013 and published the first millipede genome. This latest study provides a firmer foundation for further basic and applied research on myriapods, which in turn can contribute to studies of biodiversity and ecology.
Prof. Jerome Hui (1st right) co-chairs the new APRU Biodiversity and Pacific Ocean Programme.
Prof. Hui added, ‘In the near future, in addition to continuing to explore the hidden biology and genomics, we need to fully understand the ecological roles in soil and forest ecosystems of this fascinating yet neglected group of organisms. Hopefully one day we can better understand these life forms on land, the effects of climate change on them and their contribution to nutrient recycling, and perhaps eventually achieve zero hunger via agricultural applications.’
Prof. Hui is Co-chair of the new Biodiversity and Pacific Ocean Programme of the Association of Pacific Rim Universities which seeks to promote collective action to address biodiversity loss, protect ecosystems and combat the impacts of climate change. Co-leading the initiative with the University of Sydney and the University of California, Davis, other members include Universidad de los Andes, University of Malaya and University of the Philippines.
Read more: CUHK builds a genome bank of myriapods offering clues to the divergent behaviours of centipedes and millipedes
July 29, 2022
more
APRU Kicks off Biodiversity Genomics Program and Network
The APRU Biodiversity Genomics Program and Network was launched with an inaugural symposium on November 30. The event brought together leading genomics experts from the region to discuss progress in this area, the challenges they face, and how collective action can advance biodiversity genomics. While ‘10–15 million eukaryotic species and perhaps trillions of bacterial and archaeal species adorn the Tree of Life, ∼2.3 million are actually known, and of those, fewer than 15,000, mostly microbes, have completed or partially sequenced genomes’ (1)
“We are all very aware of threats to the world’s biodiversity so APRU has developed this symposium in biodiversity genomics to share best practices and discuss challenges. We are doing this in partnership with the University of Sydney, The Chinese University of Hong Kong and UC Davis to share best practices and discuss challenges in biodiversity genomics, which we hope will lead to the development of a region wide program supporting important capacity building activities,” APRU Secretary General Christopher Tremewan said, emphasizing the importance the program will have in addressing the biodiversity challenges of the region.
Nathan Lo, Professor of Evolutionary Biology at the University of Sydney, together with Jerome Hui, Associate Professor at The Chinese University of Hong Kong’s School of Life Sciences, led on the development of the Symposium and moderated the event.
Harris Lewin, Professor of Evolution and Ecology at the University of California, Davis, gave the keynote address on the Earth BioGenome Project, an initiative that aims to sequence and catalog the genomes of all of Earth’s currently described eukaryotic species over a period of ten years. Lewin warned that eukaryotic life is under threat from pollution, over-exploitation, invasive species and, even more alarmingly, climate change.
Andrew Jackson Crawford, Associate Professor at the Department of Biological Sciences of the Universidad de los Andes, presented on challenges and solutions for biodiversity genomics in Colombia. Dr. Carolyn Hogg, Senior Research Manager for the Australasian Wildlife Genomics Group in the Faculty of Science of The University of Sydney, explained how genomics could be used to reduce the rate of species extinction in Australia. The other speakers were Dr. Herawati Sudoyo, Deputy Director of the Eijkman Institute for Molecular Biology in Jakarta; Balaji Chattopadhyay, Assistant Professor at the Trivedi School of Biosciences at Ashoka University; Subha Bhassu, Professor of Animal Genetics and Evolutionary Biology at the University of Malaya’s Institute of Biological Sciences; Hayde Galvez, Assistant Professor and Researcher at the University of the Philippines Los Banos; and program co-leader Jerome Hui.
Following the presentations, Kathy Belov, Pro-Vice Chancellor (Global Engagement) and Professor of Comparative Genomics at the University of Sydney, explained the APRU Biodiversity Genomics Program and Network’s future agenda, encouraging anyone interested to join the group to approach the organizers.
“Over the coming months we will hold a series of workshops with the aim of building a new APRU strategic project focused on Biodiversity. This platform will give biodiversity researchers around the Pacific Rim an opportunity to join together to tackle biodiversity decline in our region using the latest advances in science. Working together we can influence policy and tackle one of the greatest challenges facing our region – the loss of our iconic native species,” Belov said.
Find out more details about the symposium here.
______________
(1) Earth BioGenome Project: Sequencing life for the future of life
Harris A. Lewin, Gene E. Robinson, W. John Kress, William J. Baker, Jonathan Coddington, Keith A. Crandall, Richard Durbin, Scott V. Edwards, Félix Forest, M. Thomas P. Gilbert, Melissa M. Goldstein, Igor V. Grigoriev, Kevin J. Hackett, David Haussler, Erich D. Jarvis, Warren E. Johnson, Aristides Patrinos, Stephen Richards, Juan Carlos Castilla-Rubio, Marie-Anne van Sluys, Pamela S. Soltis, Xun Xu, Huanming Yang, Guojie Zhang
Proceedings of the National Academy of Sciences Apr 2018, 115 (17) 4325-4333; DOI: 10.1073/pnas.1720115115
December 20, 2021
more
Human Development Forum Publishes A Better World Vol. 6 with APRU Contribution
Read the book now >>
For your interest the APRU report starts here>>
APRU is pleased to note that the Human Development Forum, an educational and research organization founded on close collaboration with UN agencies, UN member states, and civil sector organizations, has published the digital edition of A Better World Vol. 6.
A Better World is a series of publications that dedicates each volume to one of the 17 SDGs. The new volume covers Goal 14 – Conserve and sustainably use the oceans, seas and marine resources for sustainable development.
APRU’s contribution draws on the Pacific Ocean Program, featuring economy-specific analysis conducted by a team of experts from the University of British Columbia and the University of Washington on the ways that all SDG goals contribute or detract from SDG 14 throughout the Pacific.
APRU recommends policymaking that analyzes the contribution that each individual SDG makes to others, as this could help prioritize SDG achievements while minimizing the chances of unrealistic expectations and avoidable side-effects.
Indeed, APRU research illustrates the complexity of SDG achievements, including by demonstrating that eliminating poverty and hunger (SDGs 1 and 2) may delay the achievement of SDG 14 in the Pacific.
“By focusing on the experience and livelihoods of people, especially those in vulnerable human habitats, the book shows the benefits of best policy and practices, and how these may develop further as we come to terms with a changing and more turbulent world,” said Sean Nicklin, the Human Development Forum’s General Coordinator.
“This innovative endeavor is a striking example of sharing respective resources to engage the many official governmental, international organizations, institutions, and professional interests in displaying the extent and variety of their efforts to make the world a better place,” he added.
A Better World Vol. 6’s key subjects are coral reefs; implementation of international law; mangroves; marine and coastal ecosystem management; marine pollution; scientific knowledge; sustainable blue economy; and sustainable fisheries. It contains fascinating contributions from researchers and organizations across the world.
A number of the supporting agencies and institutions have asked to incorporate the book in their social media campaigns, including the contributing UN agencies. The Human Development Forum plans to publish the print volume in June 2020.
June 15, 2020
more
Biodiversity Essential to APEC Economies
2020 APEC ASPIRE Prize
Now accepting nominations
More information at: https://www.apec.org/aspire/aspire2020
The APEC Science Prize for Innovation, Research and Education (“ASPIRE”) is an annual award which recognizes young scientists who have demonstrated a commitment to both excellence in scientific research, as evidenced by scholarly publication and cooperation with scientists from other APEC member economies.
The ASPIRE Prize supports APEC’s mission to:
strengthen international science and technology networks;
enhance economic growth, trade and investment opportunities in harmony with sustainable development, through policies, innovative R&D and technologies, and knowledge sharing; and
improve linkages and efficiency between research and innovation.
ASPIRE 2020: BIODIVERSITY FOR A PROSPEROUS ECONOMY
Each year the APEC host economy is asked to provide a theme to guide nominations for the ASPIRE Prize to be awarded in their host year. For its host year of 2020, Malaysia selects “Biodiversity for a Prosperous Economy” as the ASPIRE nominating theme. Biodiversity is foundational for human health as it underpins the functioning of our ecosystems and dependence for food and water, climate, floods and disease, and more. This theme focuses on scientists’ contributions to biodiversity for prosperous economies across the APEC region by spurring research that contributes to local livelihoods, both traditional and modern medicines, and economic development.
Each member economy, through its representative on the APEC Policy Partnership for Science, Technology and Innovation (PPSTI), is invited to nominate one young scientist under the age of 40 to be considered for the 2020 ASPIRE Prize. Nominees should demonstrate a commitment to excellence in scientific research and cooperation with scientists from other APEC member economies in subjects such as: biology, chemistry, environmental science, physics, and other relevant fields.
ELIGIBILITY
Any citizen of an APEC member economy is eligible to be nominated for the ASPIRE Prize. He/she must be living at the time of his/her nomination and be under the age of 40 as of 31 December of that year (i.e., all 2020 nominees must be under the age of 40 as of 31 December 2020).
SELECTION PROCESS
Each member economy, through its representative on the APEC Policy Partnership for Science, Technology and Innovation (PPSTI), is invited to nominate one young scientist under the age of 40 to be considered for the ASPIRE Prize.
Individually qualified applicants are encouraged to complete the “Local Nomination Form” and send it to PPSTI Program Director Ms. Eva Nakamura ([email protected]) by 15 May 2020 so it may be directed toward local economy reviewers.
Once nominations are received, PPSTI members rank the nominees through a selection ballot to determine the winner. PPSTI members are asked to judge the nominees based on how well they have demonstrated:
Excellence in scientific research, as evidenced through scholarly publication;
Commitment to cooperation with scientists from other APEC member economies; and
Contribution to the theme of “Biodiversity for a Prosperous Economy.”
The winner will be recognized at an award ceremony during the 16th APEC PPSTI Meeting in Malaysia tentatively scheduled for August 2020.
ASPIRE PRIZE SPONSORS
Wiley and Elsevier, two of the world’s leading publishers of scholarly scientific knowledge, have committed to funding prize money in the amount of $25,000 USD.
February 17, 2020
more
A Proposal to Address Marine Debris and Microplastics
APRU academic experts informed APEC officials at a workshop hosted by the APEC Marine Sustainable Development Center on Marine Debris and Microplastics held in Xiamen, China, Dec 3-5, 2019.
The discussion on governance of the Pacific Ocean and the affiliated fight against microplastics and marine debris has become dramatically important, given that plastic waste has been continuously impacting the marine eco-system, inflicting tremendous damage on coastal communities’ livelihood and posing a great threat to sustainable marine development.
APRU’s Deo Florence Onda, an assistant professor at the University of the Philippines Diliman’s Microbal Oceanography Laboratory, spoke on the role of microbes on the fate of plastics in the marine environment, and APRU’s Christelle Not, an assistant professor at the University of Hong Kong’s Department of Earth Sciences, presented her evaluation of the plastic pollution in Hong Kong and its link with the global plastic issue.
“Although we are still at the early stage of monitoring the level of plastic pollution in Hong Kong waters, we are already observing a strong spatial and temporal variability in the abundance of microplastics,” Not said.
“It is clear that this is impacting the marine ecosystem and we found that fish and crabs from Hong Kong have ingested microplastics,” she added.
The workshop was organized by the APEC Marine Sustainable Development Center and Third Institute of Oceanography, with more than 120 representatives from 11 APEC economies taking the opportunity to exchange ideas on policy, scientific research, and the public and private sectors’ participation in addressing marine debris and microplastics.
APRU contributed to the development of the Blue Citizen Partnership Initiative conceptualized to engage those who are willing to learn about the ocean and to practically involve themselves in marine environment protection.
The initiative is designed to heighten citizens’ awareness of reducing marine debris, to develop their concept of sustainable marine development, and to deepen their understanding of all the dangers that are associated with marine debris and microplastics pollution.
It aims to accelerate investment in education of marine science and scientific knowledge to address waste sorting; encourage the “greening” of industries; and promote eco-wise behaviours and lifestyles, including by motivating tourists to resolutely cut down their use of disposable plastic products. The Blue Citizens Initiative will continue to be developed within APEC.
A clean ocean is a common vision, APRU is supportive of the Blue Citizens Initiative and through the APRU Pacific Ocean Program joins the relentless pursuit of protecting the global marine community.
January 20, 2020
more
What are the co-benefits to SDG14 when making progress toward other SDGs? Initial findings reported at APEC SOM3 from the APRU Pacific Ocean Program
Leading marine science expert of APRU’s Pacific Ocean Program on advancing UN Sustainable Development Goal (SDG) 14: Life Below Water informed policymakers on early findings of the program at the Third Senior Officials’ Meeting (SOM3) in Puerto Varas, Chile, in August.
APRU’s Inaugural Pacific Ocean Cluster Project: Advancing SDG 14 for the sustainable future of the Pacific Ocean focuses on enhancing sustainable development of coastal states, communities, and economies around the Pacific-Rim region. The overall aim is to provide policy pathways to advance SDG 14.
A team of experts from The University of British Columbia and University of Washington have conducted economy-specific analysis of the ways that all SDG goals contribute or detract from SDG 14 throughout the Pacific, with the initial results indicating a potential asymmetry in SDG alignment and achievements.
From this team, Gerald Singh, now an assistant professor at the Department of Geography of the Memorial University of Newfoundland indicates that these initial results means that while making progress to achieve SDG 14 there are benefits to SDGs 1 and 2 of ending poverty and hunger (though not fully achieve these goals). However, fully achieving the goals of eliminating poverty and hunger by the 2020-2030 achievement dates may prevent the achievement of SDG 14 in the Pacific.
Singh furthermore explained that the achievement of the SDG 14 in the Pacific is also being complicated by the economies not clustering according to classic development categories such as “developed”, “developing”, and “transitioning” but instead including a mix of fully developed and developed economies.
In view of these findings, it is the project team’s key objective to collaborate and explore ideas with the OFWG [APEC’s Oceans and Fisheries Working Group] more closely.
“One area for collaboration can be through data sharing across projects to support comparison and verifying project results,” he added.
Singh’s presentation to APEC OFWG and initiated and supported through the APRU Pacific Ocean Program generated great interest by some member economies as well as non-member guests.
Next steps included discussions of the possibility of future collaboration with the delegations of China; the Philippines; the Coral Triangle Initiative on Coral Reefs, Fisheries, and Food Security; the Ocean Conservation Administration Ocean Affairs Council (in Chinese Taipei); as well as The Nature Conservancy.
The SOM3 is the last senior officials’ preparatory meeting before the APEC Economic Leaders’ Meeting (AELM) in November.
Held under the theme “Connecting people, building the future,” it facilitated fruitful discussion surrounding the priority areas of digital economy, regional economic integration, connectivity, marine cooperation, and women and inclusive growth.
August 22, 2019
more
The APEC 2018 Workshop on Innovative Marine Debris Solutions, July 26-27, 2018, Beijing
The issue of marine debris has received high attention from economies, international organizations and multiple fora.
The Workshop on Best Practices Sharing on Marine Debris Management in Coastal Cities of APEC Region was held in Xiamen on Nov 4-5th, 2017. The workshop outputs were put into the United Nations General Assembly Resolution on “Oceans and the law of the Sea” as Article 215.
This forthcoming workshop taking place July 26-27, 2018 proposes to collect innovative approaches and to share best practice to address marine debris in the APEC region. Click here to see the proposal.
The workshop objectives are to:
1) collect innovative approaches addressing marine debris;
2) share best practice, information, and technologies to reduce marine debris in the APEC region;
3) encourage and promote Public Private Partnerships.
The event, hosted in partnership with Peking University, is aimed at managers/policy makers, researchers, and private-sector participants and will feature a 1-day meeting and 1-day scientific tour.
The APEC Marine Sustainable Development Center China is making funding available for one APRU scholar to contribute to the session addressing new research advances on marine debris and micro-plastics.
See a post-event report from Peking University here.
July 3, 2018
more
Call for APRU Expert Engagement for the development of the APEC Marine Sustainable Development Report 2
The first APEC Marine Sustainable Development Report (AMSD) developed by the APEC Marine Sustainable Development Centre received endorsement in 2014. The AMSD is the first comprehensive report in APEC to research the status and progresses of marine sustainable development of the Asia-Pacific region. Download the AMSD here
At the 2016 APEC Ministerial Meeting in Peru it was proposed to update the AMSD to promote regional marine sustainable development as a key APEC contribution to the 2030 Agenda for Sustainable Development.
The proposed theme of report 2 is ‘APEC Sustainable Development Report: Sustainable Development Goals in APEC’.
The report objectives are to:
1. reflect trends and endeavors of APEC and its member economies in achieving SDGs, especially SDG 14 and other goals and objectives relevant to ocean and coasts;
2. map and take stock of OFWG’s projects and activities relevant to marine sustainable development;
3. serve as a platform to take APEC’s active role in facilitating the implementation of 2030 agenda in Asia-Pacific region.
The APEC Marine Sustainable Development Centre is currently bringing together a core group of experts to support the development of the general report (which will be supplemented by a collection of economic reports) and is calling for nominations from APRU member experts with the following backgrounds:
1. marine management and policy especially marine pollution control
2. marine ecosystem conversation and resource management
3. ocean and climate change and sustainable fisheries management
Roles and Responsibilities of the core expert group
a) develop the general report by setting outline, collecting useful data and information, drafting and reviewing;
b) have sufficient communication during the formulation process of the report through workshops, informal meetings, tele-meetings and emails;
c) work in collaboration with APEC Marine Sustainable Development Centre to finalize and publish the final report;
d) design the questionnaire to collect relevant information from member economies for the purpose of drafting AMSD Report 2.
e) work on other issues concerning the developing and updating of the report;
A 2-day workshop is expected to be convened in Xiamen sometime between July and September 2017 for shaping the theme, chapter structure and outlines of the report. Date of the workshop is to be determined after the establishment of core expert group. Some funding maybe available to support academic expert participation. TBC in due course.
See the information sheet for more detail about background, proposed work plan, methodologies and time frame of overall project.
Nomination form for the core expert group will need to be submitted to [email protected] via emailed by end of Friday, May 26, 2017.
APRU will forward suitable nominations to the APEC project team who will contact selected experts by the middle of June 2017.
See here for more information about the APEC Marine Sustainable Development Centre
For more information about the report, please contact [email protected]
For more information about the application process, please email to [email protected]
Download attachments:
Information_Sheet_concerning_Nomination_of_the_AMSD_report.pdf
APRU_Experts_Recommendation_Form_of_AMSD_report.docx
May 22, 2017
more